Workshop report: Developmental Biases in Evolution
20 November 2018
“Developmental bias is a manifestation of a much more fundamental principle, and is the norm rather than the exception”, according to one scientist. “Developmental bias is a misleading term and we should get rid of it”, according to another. Both were in the same room, at the same time. So what is developmental bias, what role does it play in evolution, and why do we even care? These questions were the focus of an interesting and spirited workshop, the third in a series of EES project meetings, held recently at the Santa Fe Institute. As with the first meeting, I was able to sit in on this workshop, like a fly on the wall, and listen to the presentations, arguments, and debates. Here’s what I learned…

Developmental bias
The classical view in evolutionary theory is that genetic mutations are purely chance events, with an equal probability of a mutation happening anywhere in the genome and resulting in any of the three possible substitutions. In other words, a nucleotide A in position 135 on chromosome 8 is equally likely to be mutated to a C as a mutation from a G to a T in position 26 on chromosome 13. Even though there is evidence that this is not always true, such a fully uniform probability distribution for genetic mutations is, to a first approximation, a reasonable assumption and this assumption is often built into population and quantitative genetics theory (e.g. as symmetrical Gaussian distributions).
This ‘uniform probability’ assumption is often carried over to the phenotypic level as well. In other words, it is assumed that all conceivable ‘mutations’ (variations) in phenotypic traits are possible and, moreover, equally likely. For example, it is assumed that animals could, in principle, develop any number of digits or segments, in any relative size proportion, or develop any kind of coloration or pigmentation, all with equal probability.
This assumption is incorrect. In fact, phenotypic variants produced in response to genetic or environmental perturbation are, in many cases, not uniformly distributed, but rather channeled or directed by the processes of development. This phenomenon, known as developmental bias, not only limits what is possible but can also facilitate evolutionary adaptation, and thus give it directionality, according to Tobias Uller of Lund University, one of the co-organizers of the workshop.
As philosopher of science Ingo Brigandt (University of Alberta) pointed out, the notion of developmental bias goes back several decades to the conceptual ideas of, among others, Riedl (1977), Gould & Lewontin (1979), Oster & Alberch (1982), and Maynard-Smith et al. (1985). Initially referred to as developmental constraints, the term developmental bias was adopted later on to emphasize that it can also facilitate evolution, besides constraining it.
Morphospaces
One way to visualize and quantify developmental bias is through the use of morphospaces. A morphospace is a graphical, multi-dimensional representation of the possible form, shape or structure of a phenotypic trait or an entire organism. In particular, each axis (dimension) in a morphospace corresponds to a variable which describes some character of the phenotypic trait or organism of interest. This method was originally introduced by Raup (1966). A simple example of a morphospace is given in the figure below.
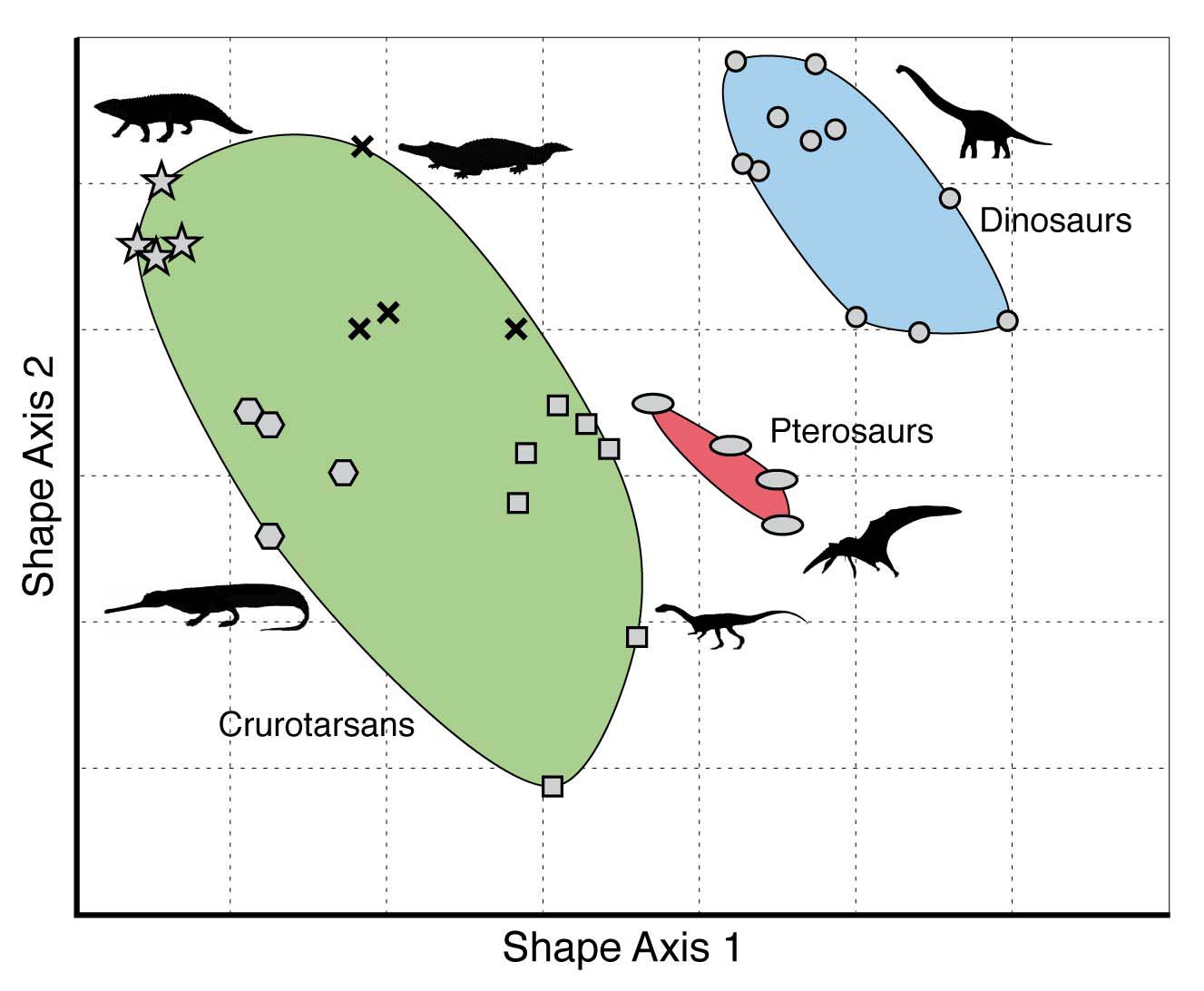
As this morphospace clearly shows, it is not filled uniformly. However, this in itself is not (yet) evidence for developmental bias. For example, it could be that one or more of these reptile groups went extinct before they had a chance to ‘explore’ (through genetic mutations giving rise to phenotypic variation) the entire morphospace. Or maybe they did explore some (now empty) parts of the morphospace, but those particular reptilian shapes were not fit enough to survive, and were therefore ‘eliminated’ through natural selection.
To investigate the presence (or absence) of developmental bias, though, we can subject organisms (or specific phenotypic traits) to experimental investigation, for instance through artificial selection, rather than relying on natural selection alone. This may be somewhat difficult to do with dinosaurs, but luckily there are plenty of more benign organisms that can be easily bred in the laboratory. One such study was done with butterflies.
Experimental evidence
Paul Brakefield (University of Cambridge) presented some wonderful selection experiments on eyespots on butterfly wings. Starting with a ‘wild type’ (i.e., specimens as they exist in the wild), and focusing on two ‘target’ eyespots, these butterflies were then bred in the laboratory, and selected for different eyespot characters. For example, one set of lineages were specifically selected for larger or smaller eyespots, while in another experiment lineages were selected for specific types of eyespot color combinations (more gold or more black). The results are presented in the morphospaces in the next figure.
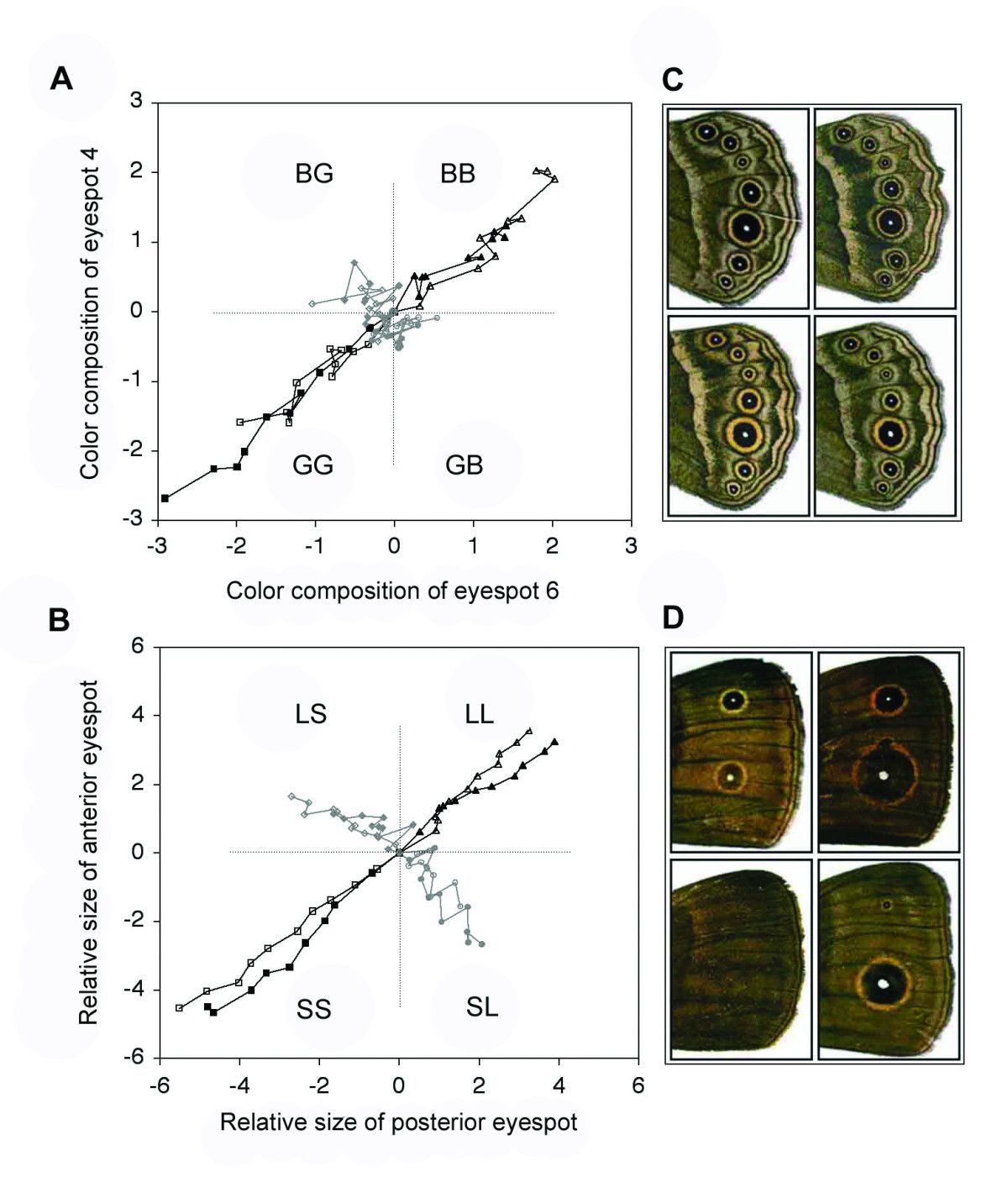
Starting from the wild type (at the center of the morphospace), relative eyespot size can be varied in all four directions, i.e., in the direction of all four corners of the morphospace. This is illustrated in image B (bottom) in the figure. However, color combinations only varied in two out of the four possible directions: either to the top-right or to the bottom left of the morphospace, as image A (top) shows.
So, while relative eyespot size variation seems to be distributed quite uniformly, color combination variation appears to have a clear bias. Somehow the butterflies are not able to develop certain eyespot color combinations (i.e. one spot more gold and the other more black), while others are developed quite easily (both spots more gold, or both more black).
Similar convincing evidence for developmental bias was presented by Kathryn Kavanagh of the University of Massachusetts, based on relative sizes of phalanges (bone segments such as the little bones in your fingers and toes). Her results are presented in the figure below, showing highly biased (i.e., non-uniform) distributions.
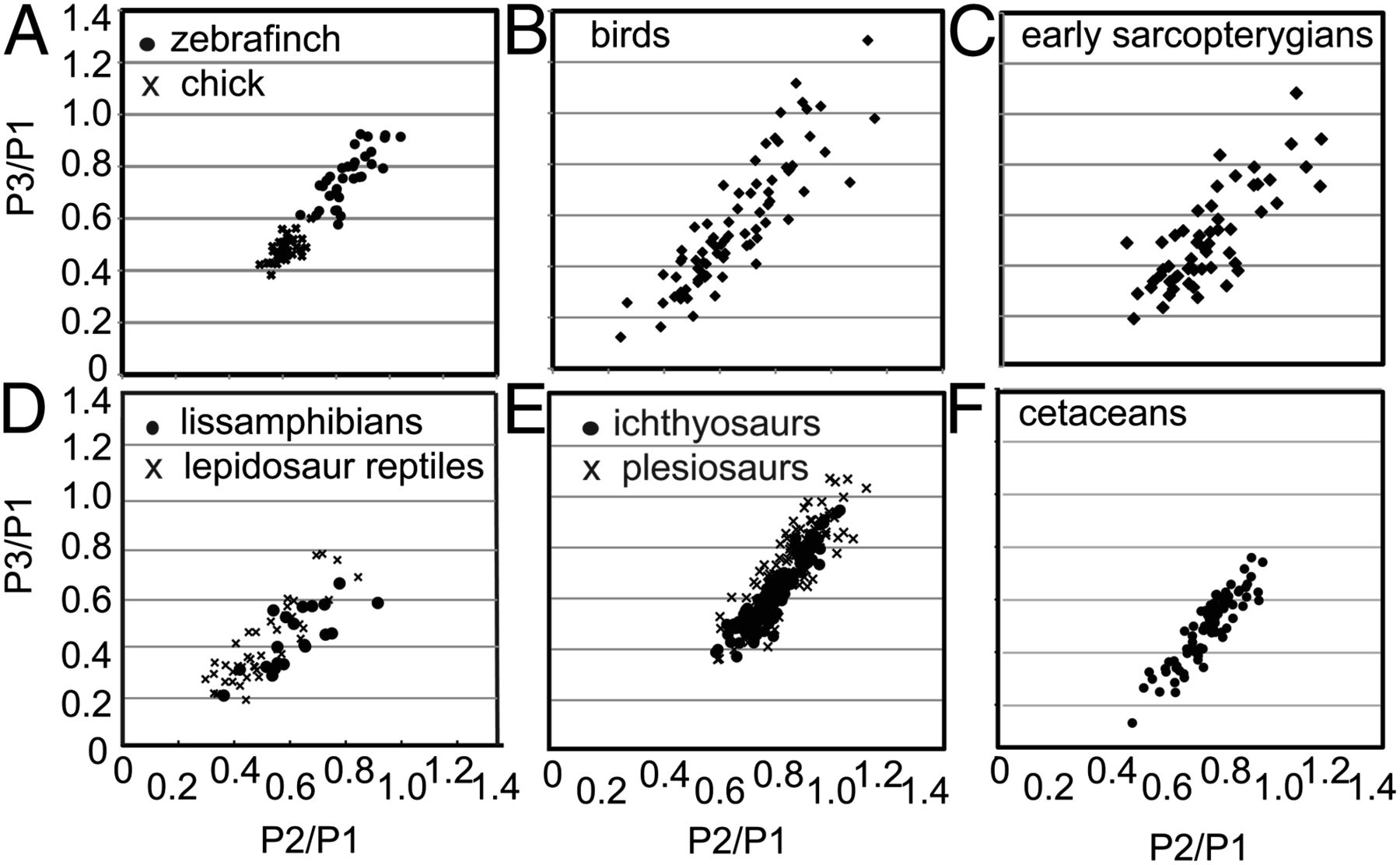
More compelling still, both Brakefield and Kavanagh showed that they could use knowledge of bias generated from laboratory experiments in model animals to predict which combinations of traits would be observed across related taxa in nature.
Directionality in evolution
One plausible explanation for such developmental bias is that the traditional ‘one gene for one trait’ view is simply wrong. Most genes influence multiple traits, and most traits are influenced by multiple genes. Thus, if you want to make a change in one phenotypic trait (through a genetic mutation), you inevitably also change one or more other traits. In other words, trait variations are correlated, and therefore not uniformly distributed.
This seems to be especially the case for traits that are determined early on in development, which generally have much lower variability than traits that are determined later on, according to Frietson Galis of the Naturalis Biodiversity Center. The more correlated (i.e., developmentally connected) traits are, the harder they are for evolution to change without disrupting other traits. Or as Anjali Goswami of the Natural History Museum put it: “When considering development in your model, convergence becomes the expectation.”
On the other hand, though, developmental bias can also facilitate evolutionary change. David Jablonski (University of Chicago) argues that evolution follows a path of developmental preference. Those phenotypic trait variations that are developmentally easier to achieve are likely to have a selective advantage over developmentally more difficult variations, even if the underlying genetic mutations required for them are equally likely. And as Doug Erwin (National Museum of Natural History) argued, although the Cambrian explosion was largely driven by environmental changes, it could only have happened due to an earlier evolutionary adaptation towards a more complex molecular regulatory system that allowed different animal groups and body sizes to develop in the first place. In other words, the trigger for the Cambrian explosion was environmental, but the facilitator was developmental.
Other examples presented include the emergence of herbivory and animal domestication. According to Scott Gilbert of Swarthmore College, an herbivory lifestyle and morphological adaptation was only possible after the symbiosis of certain animals and particular gut-bacteria that are able to digest cellulose. More generally, he claims, symbionts scaffold animal development and evolution. And Adam Wilkins of the Humboldt University presented the case of ‘domestication syndrome’, already recognized by Darwin, for which a set of seemingly unrelated traits (such as smaller brain size, curly tails, and piebald pigmentation in animals) come together with selection purely for tameness, most likely because of a connection between the nervous and endocrine systems.
Theoretical considerations
All these examples form an impressive body of evidence for developmental bias and its possible influence on the course of evolution. According to David Krakauer, president of the Santa Fe Institute, developmental bias is to be expected due to the grounding of biology in physical space and matter. In particular, many-to-one mappings (such as many genes influencing a single trait) “make forms few and similar; regulation constrains developmental variability.” He illustrates his point with an extreme example of “life existing along just one dimension”.
A famous scaling law in biology states that metabolic rate scales with body mass to the power of 3/4. The figure below illustrates this for a wide range of organisms over several orders of magnitude in body mass. This figure can be interpreted as a morphospace; one with an extremely non-uniform distribution of variation. Indeed, all observations fall almost perfectly along a straight line. Of course one could argue that it is purely natural selection that forces these points to fall along a straight line. But there are theoretical arguments that this is caused at least in part due to physical forces imposing constraints, or bias.
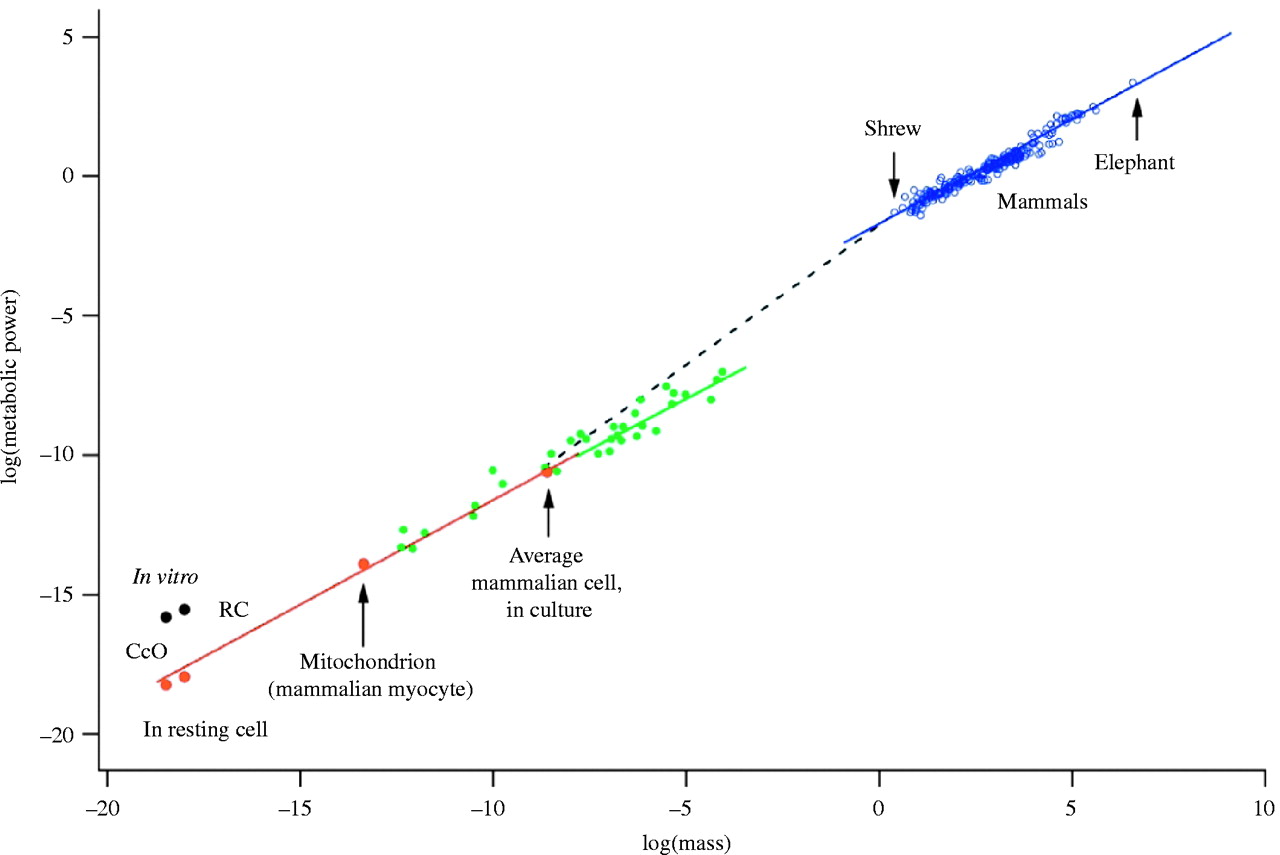
West, Brown & Enquist (1997) showed that this scaling law can be explained mathematically by taking three-dimensional space into account, and the need for a space-filling, fractal-like vascular system to deliver nutrients to all parts of an organism’s body. From this mathematical argument, the scaling exponent of 3/4 follows automatically. In other words, organisms cannot have just any form or shape, but they are (developmentally) constrained by this scaling law. Hence Krakauer’s argument that a strong developmental bias is to be expected in biology, due to its grounding in physical space and matter. A computational model developed by Borenstein and Krakauer (2008) showed how regulation rapidly evolves in developmental systems, such that bias is inevitable.
However, if a bias is to be expected, is it still a bias? After all, a bias is only a bias to the extent that it deviates from your expectation. But if a deviation is to be expected in the first place, then why call it a bias? Any developmental bias, like those presented at the workshop, is only a bias if your expectation is that of a uniform distribution. But this is exactly the standard view of evolution that proponents of the extended evolutionary synthesis are trying to criticize! As Isaac Salazar-Ciudad of the Autonomous University of Barcelona pointed out quite passionately, this is logically inconsistent, and the term ‘developmental bias’ should therefore not be used at all.
I have to say that I cannot disagree with this argument. Even workshop co-organizer Tobias Uller reluctantly admitted that maybe in the long run it is indeed not the right term, but for now it is a useful vehicle for stimulating debate and making scientific progress.
Conclusions
Call it bias or not, the many examples presented at the workshop (only some of which are mentioned here) make it clear that there is a highly non-uniform distribution of variability in biological organisms, largely due to their developmental processes. Several of the examples also make it clear that this has direct consequences for the course of evolution, possibly resulting in directionality in evolutionary change, in particular along a path of developmental preference. This certainly seems to challenge the standard view of uniform variability, and natural selection being the only source of directionality in evolution.
As with the first EES project meeting, I thoroughly enjoyed being a (hopefully unbiased) fly on the wall during this workshop, and learned many new and exciting things about evolution. It remains worthwhile to continue following these debates and new developments in evolutionary biology.
